- About
-
Solutions
-
Services
-
Biosciences
- PureDisc™ Enabled Membrane Protein Production
- Assay Development and Transfer
- Biochemistry / Enzymology
- Biomarker Services
- Biophysics
- Cellular Biology
- Compound screening and cascade design
- Cryo-EM
- Fibrosis Assays
- Project Management and Consultancy Services
- Protein Expression
- Structural Biology
- X-ray Crystallography
- Chemistry
- Integrated Drug Discovery
- Computer Aided Drug Design
- Hit ID and Screening Services
- DMPK Services
-
Biosciences
- Target Classes and Modalities
- Therapeutic Areas
-
A-Z
- A
- B
- C
- D
- E
- F
- G
- H
- I
- K
- L
- M
- N
- O
- P
- R
- S
- T
- V
- X
-
Services
- Library
- News & Events
- Careers
- PureDisc™
Protein-Protein Interactions
Identifying small molecules that disrupt PPIs
Protein-protein interactions (PPIs) are vital to the regulation of biological systems, and PPI dysfunction is often implicated in the development of disease. However, the identification of small, drug-like molecules which disrupt the interaction between two proteins can be challenging.
High-resolution structures show that some PPIs occur between relatively large, featureless protein surfaces, and this is a distinct contrast to the deep, well-defined pockets offered by enzyme targets (such as kinases or proteases). However increasing numbers of PPIs are being shown to have druggable pockets or ‘hotspots’, where amino acid side chains, or an element of the secondary structure such as an α-helix, mediate the interaction.1,2 These features offer opportunities for drug design, and a handful of PPI inhibitors have now reached the market (e.g. Tirofiban and Maraviroc) with several more currently in clinical development.
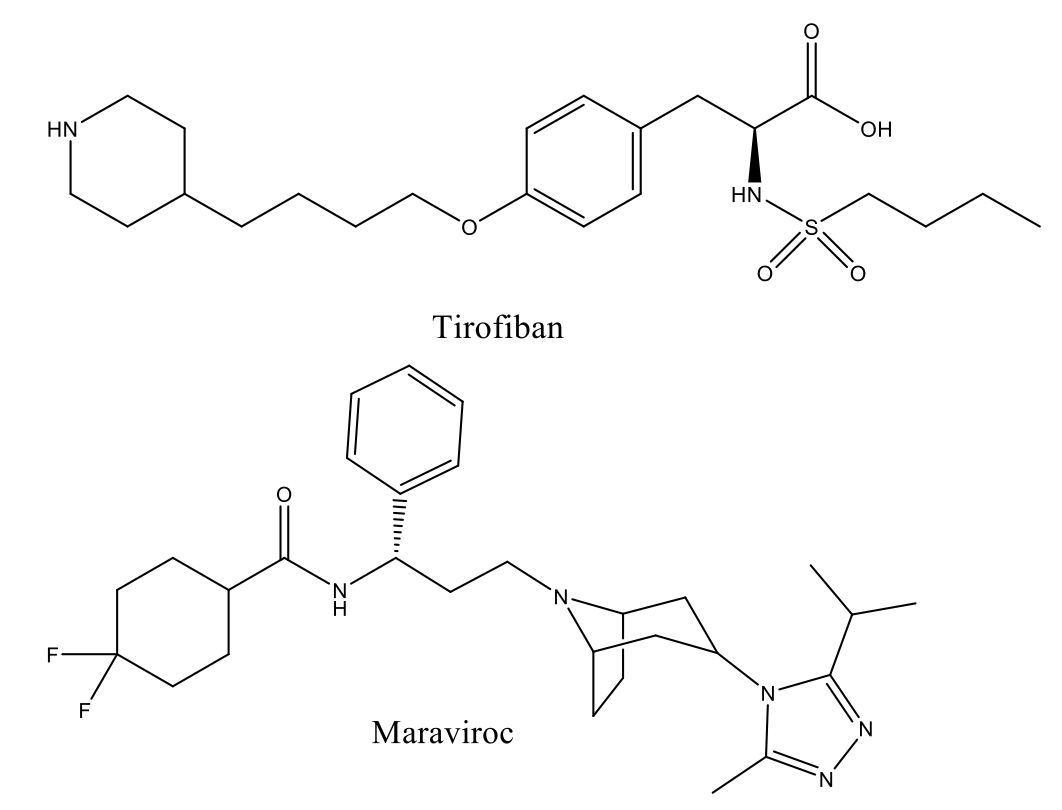
PPI Drug Discovery at Domainex
Domainex scientists have successfully worked on the design of novel inhibitors for several PPIs, for both known and innovative drug targets. We have deployed a variety of medicinal chemistry strategies to successfully crack these difficult targets. For example, Domainex worked with Ark Therapeutics on the design of antagonists of the PPI between neuropilin-1 and vascular endothelial growth factor-A (VEGF-A) and employed a peptidomimetic strategy.3-5 Click here for the full case study.
Expression of protein constructs that are suitable for the study of PPIs can be problematic. Our proprietary protein expression technology, Combinatorial Domain Hunting, has been adapted to identify the respective regions of two proteins that are required to mediate their interaction in a variant that we call CDH2. This approach affords orthogonally tagged protein sequences that are ideally suited to the development of PPI assays.6
Our assay biologists have established a number of PPI assays using both biochemical (AlphaLISA®, FRET and fluorescence polarisation) and biophysical techniques. Cell-based assays to study PPIs can also be developed using, for example, the Promega Nanoluc®-derived BRET or HiBiT/LgBiT systems.
FLIP-FADD Inhibitors
An example of an integrated project that Domainex has work on is a collaboration with Queen’s University of Belfast to find selective inhibitors of the FLIP-FADD interaction. Domainex successfully identified hit molecules using our virtual screening platform, LeadBuilder, and the subsequent Lead Optimisation programme led to the discovery of novel, first-in-class inhibitors of this PPI. Click here for the full case study.
RAS Inhibitors
Another example is a collaboration with the University of Oxford to identify small molecules which disrupt the PPI between RAS and its effector proteins. Domainex deployed a strategy of structure-based drug design to optimise a RAS-binding fragment hit. We also developed an assay to characterise compounds that block the interaction between RAS and downstream effector proteins. The programme resulted in a series of potent, cell-permeable inhibitors which prevent RAS-effector interactions and inhibit RAS-dependent signalling in cells.7 Click here for the full case study.
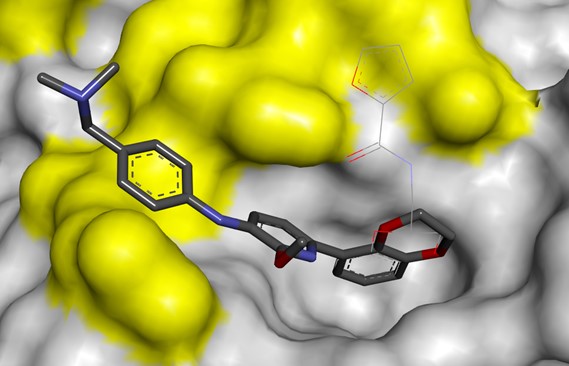
PoPPI Consortium
Domainex is an industrial member of the EPSRC-funded PoPPI (Perturbation of Protein-Protein Interactions) consortium between the Universities of Leeds and Bristol, AstraZeneca and the Northern Institute for Cancer Research, which focuses on the development of computational and chemical tools to interrogate PPIs and the development of generic insights that inform research against this class of drug targets.
If you would like to access the Domainex expertise in the field of PPIs to support your own programme we would be delighted to hear from you.
References
1. Small-molecule inhibitors of protein-protein interactions: progressing towards the dream. Arkin, M.R., and Wells, J.A. Nat. Rev. Drug Discov., 2004, 3, 301–317.
2. A hot spot of binding energy in a hormonereceptor interface. Clackson, T., and Wells, J.A. Science, 1995, 267, 383–386.
3. Small molecule inhibitors of the neuropilin-1 vascular endothelial growth factor A (VEGF-A) interaction. Ashley Jarvis, Charles K. Allerston, Haiyan Jia, Birger Herzog, Acely Garza-Garcia, Natalie Winfield, Katie Ellard, Rehan Aqil, Rosemary Lynch, Chris Chapman, Basil Hartzoulakis, James Nally, Mark Stewart, Lili Cheng, Malini Menon, Michelle Tickner, Snezana Djordjevic, Paul C. Driscoll, Ian Zachary and David L. Selwood., J. Med. Chem. 2010, 53, 5, 2215-2226.
4. N-Terminal Modification of VEGF-A C Terminus-Derived Peptides Delineates Structural Features Involved in Neuropilin-1 Binding and Functional Activity
Haiyan Jia, Rehan Aqil, Lili Cheng, Chris Chapman, Shaheda Shaikh, Ashley Jarvis, A. W. Edith Chan, Basil Hartzoulakis, Ian M. Evans, Antonina Frolov, John Martin, Paul Frankel, Snezana Djordevic, Ian C. Zachary and David L. Selwood., ChemBioChem., 2014, 15, 8, 1161-1170.
5. Small Molecule Neuropilin-1 Antagonists Combine Antiangiogenic and Antitumor Activity with Immune Modulation through Reduction of Transforming Growth Factor Beta (TGFβ) Production in Regulatory T-Cells. Jonathan Powell, Filipa Mota, David Steadman, Christelle Soudy, Jeremy T. Miyauchi, Stuart Crosby, Ashley Jarvis, Tifelle Reisinger, Natalie Winfield, Graham Evans, Aled Finniear, Tamas Yelland, Yi-Tai Chou, A.W. Edith Chan, Andrew O’Leary, Lili Cheng, Dan Liu, Constantina Fotinou, Carla Milagre, John F. Martin, Haiyan Jia, Paul Frankel, Snezana Djordjevic, Stella E. Tsirka, Ian C Zachary and David L. Selwood., J. Med. Chem. 2018, 61, 9, 4135-4154.
6. A combinatorial method to enable detailed investigation of protein-protein interactions. Kate Maclagan, Rita Tommasi, Emmanuelle Laurine, Chrisostomos Prodromou, Paul C Driscoll, Laurence H Pearl, Stefanie Reich and Renos Savva., Future Med. Chem. 2011, 3, (3), 271-82.
7. Small molecule inhibitors of RAS-effector protein interactions derived using an intracellular antibody fragment. Camilo E. Quevedo, Abimael Cruz-Migoni, Nicolas Bery, Ami Miller, Tomoyuki Tanaka, Donna Petch, Carole J.R. Bataille, Lydia Y.W. Lee, Phillip S. Fallon, Hanna Tulmin, Matthias T. Ehebauer, Narcis Fernandez-Fuentes, Angela J. Russell, Stephen B. Carr, Simon E.V. Phillips and Terence H. Rabbitts., Nat Commun 9, 3169 (2018).
Domainex press releases in the PPI field
Novel first-in-class FLIP inhibitors for the treatment of cancer
28th March 2019
A new class of RAS-effector protein-protein interaction inhibitors
20th December 2018
Queen’s University Belfast and Domainex Win Late-Stage Award to Progress Novel Lung Cancer Drug Candidate into the Clinic
26th April 2016
Queen’s University Belfast and Domainex in Cutting-Edge Collaboration to Advance Novel Lung Cancer Drug Research
20th April 2015
Case studies
Start your next project with Domainex
Contact one of our experts today