- About
-
Services
-
Offerings
- Offerings
- ADME and Bioanalytical Sciences
- Analytical Chemistry
- Assay Development
- Biochemical Assays
- Biophysical Assays
- Cell Based Assays
- Computational Chemistry
- Fragment and Compound Screening
- Integrated Drug Discovery Services
- Medicinal Chemistry
- Project Management and Consultancy Services
- Protein Expression and Purification Services
- Structural Biology
- Synthetic Chemistry
- Virtual screening
-
Research Phases
- Research Phases
- Hit Identification
- Hit to Lead
- Lead Optimisation
- Therapeutic Areas
- Target Classes
-
Approaches & Techniques
- Approaches & Techniques
- CDH (Target Gene Fragmentation)
- Cryogenic Electron Microscopy (Cryo-EM)
- Differential Scanning Fluorimetry (DSF) and nanoDSF Services
- Direct-to-Biology (D2B)
- Dynamic Light Scattering (DLS)
- eProtein Discovery
- Flow Cytometry
- Fragment Based Drug Discovery (FBDD)
- FragmentBuilder
- Grating-Coupled Interferometry
- High Throughput Screening
- Isothermal Titration Calorimetry (ITC)
- LeadBuilder
- Photoaffinity Labelling
- PoLiPa (Membrane Protein Solubilisation)
- Spectral Shift and MST Services
- Structure Based Drug Design (SBDD)
- Surface Plasmon Resonance (SPR)
- X-ray Crystallography
-
Offerings
- Library
- News & Events
- Careers
G9a: Lysine Methyltransferase Inhibitors for the Treatment of Cancer
Exciting lead compounds for treatment of a range of solid tumours
Domainex has solved the key technical challenges associated with KMTs, including generating a number of turnover assays and proprietary crystal structures.
Challenge
Lysine methyltransferases (KMTs) are involved in epigenetic gene regulation. Histones are proteins around which the DNA is wrapped in chromatin, and covalent modification of the histone amino-acid side chains generates an epigenetic ‘code’ that determines whether the associated gene is expressed or repressed. KMTs catalyse the transfer of methyl groups from S-adenosyl methionine (SAM) to lysine residues on histone proteins (Figure 1). G9a mono- or di-methylates Histone 3 at Lysine residue 9 (H3K9) and this represses gene expression. Literature supports the role of G9a in mechanisms of carcinogenesis, making it an attractive oncology target.1-5
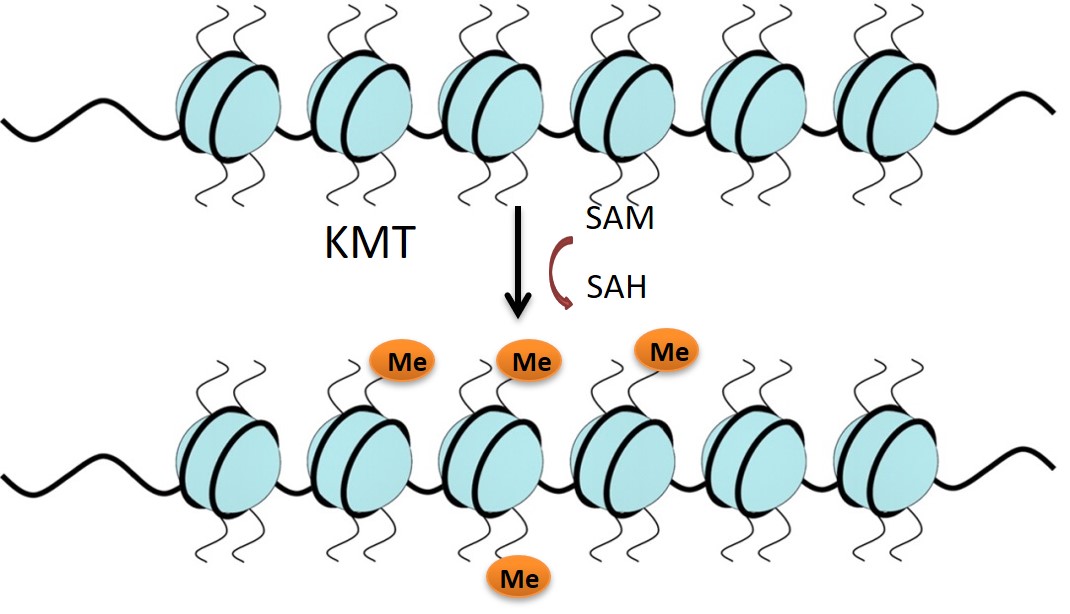
Solution
Hit finding against this challenging target used the Domainex proprietary virtual screening platform, LeadBuilder. The virtual screening strategy for this target was designed using an in-house crystal structure of a G9a-substrate peptide complex, as well as some published information. About 1200 compounds were selected and tested in an AlphaScreen biochemical assay devised by our Assay Biology team which resulted in the identification of eight confirmed hits. One series was taken forward into a hit to lead programme and, after the design and synthesis of just 200 compounds by Domainex’s computational and medicinal chemists, exciting lead compounds were identified, e.g. DMX8.1.
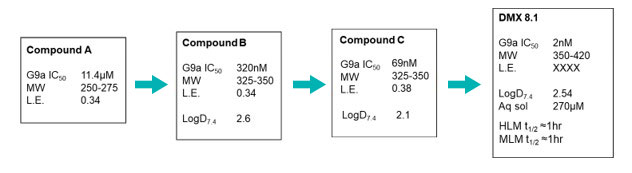
DMX8.1 is a potent inhibitor of G9a (IC50 2nM) with molecular and physical properties that are favourable for optimisation into oral drugs. As we intended, the series is non-SAM competitive and binds in the substrate-binding groove (see Figure 3). The lead molecules are novel with a completely distinct chemical scaffold from reported G9a/GLP inhibitors (e.g. BIX-012946 and UNC compounds7,8, BRD47709 and A-36610).
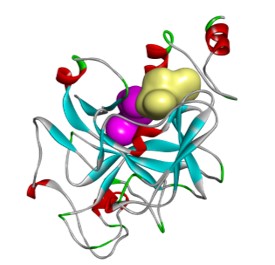
A collaboration has been forged with QIMR Berghofer that combines QIMR Berghofer’s expertise in KMT biology with Domainex’s expertise in medicinal chemistry, biochemistry, biophysics and structural biology. This collaboration has allowed the lead compounds to be further characterised.

We have shown that DMX8.1 does not reduce global levels of H3K9 methylation and has an effect at specific G9a-regulated promoters (e.g. LC3B; see Figure 4).
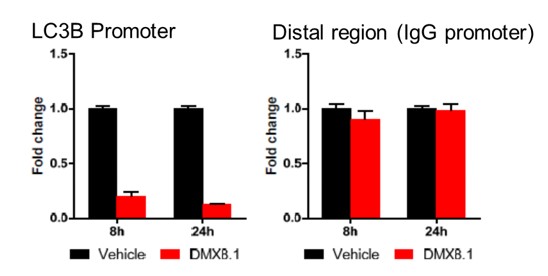
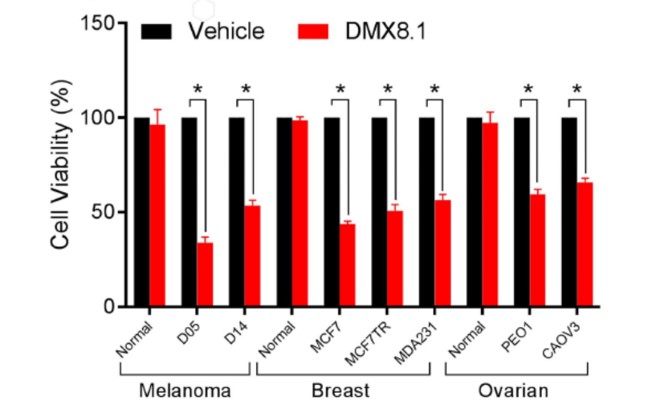
DMX8.1 was also shown to reduce the viability of malignant cell lines in an MTT survival assay (Figure 5), whilst leaving ‘normal’ cell-lines unaffected.
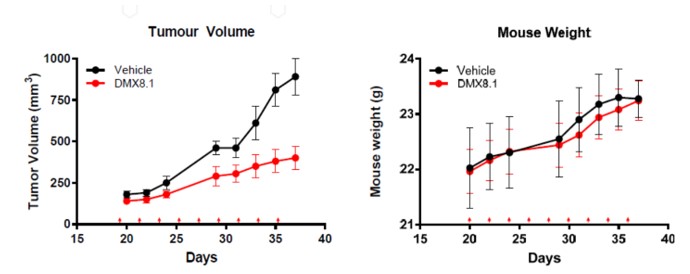
Next DMX8.1 was assessed in a triple negative breast cancer tumour xenograft mouse model (Figure 6). It significantly reduced the tumour volume while having no statistically relevant effect on the weight of the mice, nor any visible signs of ill health.
Conclusions
Starting from a LeadBuilder virtual screen we have identified novel potent inhibitors of the lysine methyltransferase G9a, which have potential to be developed as treatments for a range of solid tumours. DMX8.1 has good metabolic stability both in vitro and in vivo (data not shown) and has demonstrated single-agent efficacy in a tumour xenograft mouse model. It has shown no adverse effects in studies to date including a PK study at 10x the efficacious dose.
Domainex Expertise
• Integrated Drug Discovery • Hit Identification • Virtual Screening via LeadBuilder • Assay Development
• Biochemistry • Biophysics • Computational Chemistry • Hit-to-lead • Medicinal and Synthetic Chemistry
• Structural Biology • X-ray Crystallography • Methyltransferase Drug Discovery • Oncology
References
1. Deregulation of histone lysine methyltransferases contributes to oncogenic transformation of human bronchoepithelial cells. Hideo Watanabe, Kenzo Soejima, Hiroyuki Yasuda, Ichiro Kawada,Ichiro Nakachi, Satoshi Yoda, Katsuhiko Naoki and Akitoshi Ishizaka. Cancer Cell Int., 2008, 8, 15
2. Downregulation of histone H3 lysine 9 methyltransferase G9a induces centrosome disruption and chromosome instability in cancer cells. Yutaka Kondo, Lanlan Shen, Saira Ahmed, Yanis Boumber, Yoshitaka Sekido, Bassem R. Haddad and Jean-Pierre J. Issa. PLoS One, 2008, 3, 4
3. G9a mediates hypoxia-dependent gene repression. Francesco Casciello, Fares Al-Ejeh, Greg Kelly, Donal J. Brennan, Shin Foong Ngiow, Arabella Young, Thomas Stoll, Karolina Windloch, Michelle M. Hill, Mark J. Smyth, Frank Gannon, Jason S. Lee. PNAS, 2017, 114, (27), 7077-7082
4. Targeting histone methyltransferase G9a inhibits growth and Wnt signaling pathway by epigenetically regulating HP1α and APC2 gene expression in non-small cell lung cancer. Keqiang Zhang, Jinhui Wang, Lu Yang, Yate-Ching Yuan, Tommy R. Tong, Jun Wu, Xinwei Yun, Melissa Bonner, Rajendra Pangeni, Zheng Liu, Tiger Yuchi, Jae Y. Kim and Dan J. Raz. Molecular Cancer, 2018, 17, 153
5. MYC Interacts with the G9a Histone Methyltransferase to Drive Transcriptional Repression and Tumorigenesis. William B. Tu, Yu-Jia Shiah, Corey Lourenco, Peter J. Mullen, Dharmendra Dingar, Cornelia Redel, Aaliya Tamachi, Wail Ba-Alawi, Ahmed Aman, Rima Al-awar, David W. Cescon, Benjamin Haibe-Kains, Cheryl H. Arrowsmith, Brian Raught, Paul C. Boutros and Linda Z. Penn. Cancer Cell, 2018, 34, 4, 579-595
6. Reversal of H3K9me2 by a small-molecule inhibitor for the G9a histone methyltransferase. Kubicek, S.; O’Sullivan, R. J.; August, E. M.; Hickey, E. R.; Zhang, Q.; Teodoro, M. L.; Rea, S.; Mechtler, K.; Kowalski, J. A.; Homon, C. A.; Kelly, T. A.; Jenuwein, T. Mol. Cell, 2007, 25, 473–481
7. Discovery of a 2,4-diamino-7-aminoalkoxyquinazoline as a potent and selective inhibitor of histone lysine methyltransferase G9a. Liu, F., Chen, X., Allali-Hassani, A., Quinn, A. M., J. Med. Chem. 2009, 52, 7950–7953
8. Optimization of Cellular Activity of G9a Inhibitors 7-Aminoalkoxy-quinazolines. Feng Liu, Dalia Barsyte-Lovejoy, Abdellah Allali-Hassani, Yunlong He, J. Martin Herold, Xin Chen, Christopher M. Yates, Stephen V. Frye, Peter J. Brown, Jing Huang, Masoud Vedadi, Cheryl H. Arrowsmith, and Jian Jin. J. Med. Chem. 2011, 54, 6139–6150
9. A small-molecule probe of the histone methyltransferase G9a induces cellular senescence in pancreatic adenocarcinoma. Yuan Y, Wang Q, Paulk J, Kubicek S, Kemp MM, Adams DJ, Shamji AF, Wagner BK, Schreiber SL. ACS Chem Biol. 2012, 7(7), 1152-1157
10. Discovery and development of potent and selective inhibitors of histone methyltransferase G9a. Sweis RF, Pliushchev M, Brown PJ, Guo J, Li F, Maag D, Petros AM, Soni NB, Tse C, Vedadi M, Michaelides MR, Chiang GG, Pappano WN, Med.Chem.Letts, 2014, 5, 205
Start your next project with Domainex
Contact one of our experts today