- About
-
Solutions
-
Services
- Biosciences
- Chemistry
- Integrated Drug Discovery
- Computer Aided Drug Design
- Hit ID and Screening Services
- DMPK Services
- Target Classes and Modalities
- Therapeutic Areas
-
A-Z
- A
- B
- C
- D
- E
- F
- G
- H
- I
- K
- L
- M
- N
- O
- P
- R
- S
- T
- V
- X
-
Services
- Library
- News & Events
- Careers
RNA Services
close collaboration • rapid delivery • flexible solutions • deep expertise
What is RNA?
RNA is often thought of as an unstructured linear biomolecule, but it is in fact structurally rich and capable of forming complex 3D architectures such as aptamers, riboswitches, hairpins, and G-Quadruplexes. Whilst even structured RNAs are highly dynamic, they can create well defined ligand binding pockets suitable for selective small molecule modulation.
Clinically validated RNA-binding small molecules are establishing RNA as a powerful new class of drug targets:
- Ribocil - translational inhibitor binding an FMN riboswitch
- Risdiplam - FDA approved splicing modulator targeting SMN2 pre-mRNA
- Zotatifin - translation regulator stabilising eIF4A-RNA complexes
Domainex’s Integrated Strategy for Discovering RNA-Binding Small Molecules
Domainex delivers a collaborative, rapid, and highly flexible RNA-focused discovery workflow that combines:
- RNA-focused multi-objective virtual screening via Pareto front selection
- High‑sensitivity biophysical characterisation using Spectral Shift and Grating-Coupled Interferometry (GCI)
This integrated approach, powered by Domainex’s structural and biophysical expertise, enables confident identification, validation, and mechanistic profiling of RNA-ligand interactions.
Computational Virtual Screening for RNA‑Ligand Discovery
Ligand-Based Drug Design (LBDD)
Domainex's ligand-based models rapidly triage chemical space to identify fragments and compounds resembling known RNA-binding chemotypes. These approaches leverage key molecular descriptors including:
- Shape complementarity
- Electrostatic similarity
- Pharmacophore similarity
- Physiochemical properties
This enables rapid exploration of RNA-relevant chemical space and prioritisation of compounds with the highest likelihood of productive RNA interactions.
Structure-Based Drug Design (SBDD)
To complement ligand-based approaches, Domainex deploy advanced structure-based workflows specifically tailored to RNA targets. These incorporate:
- RNA pocket detection algorithms
- State-of-the-art RNA-ligand docking
- AI-driven RNA binding probability models
These approaches can identify and characterise druggable binding sites
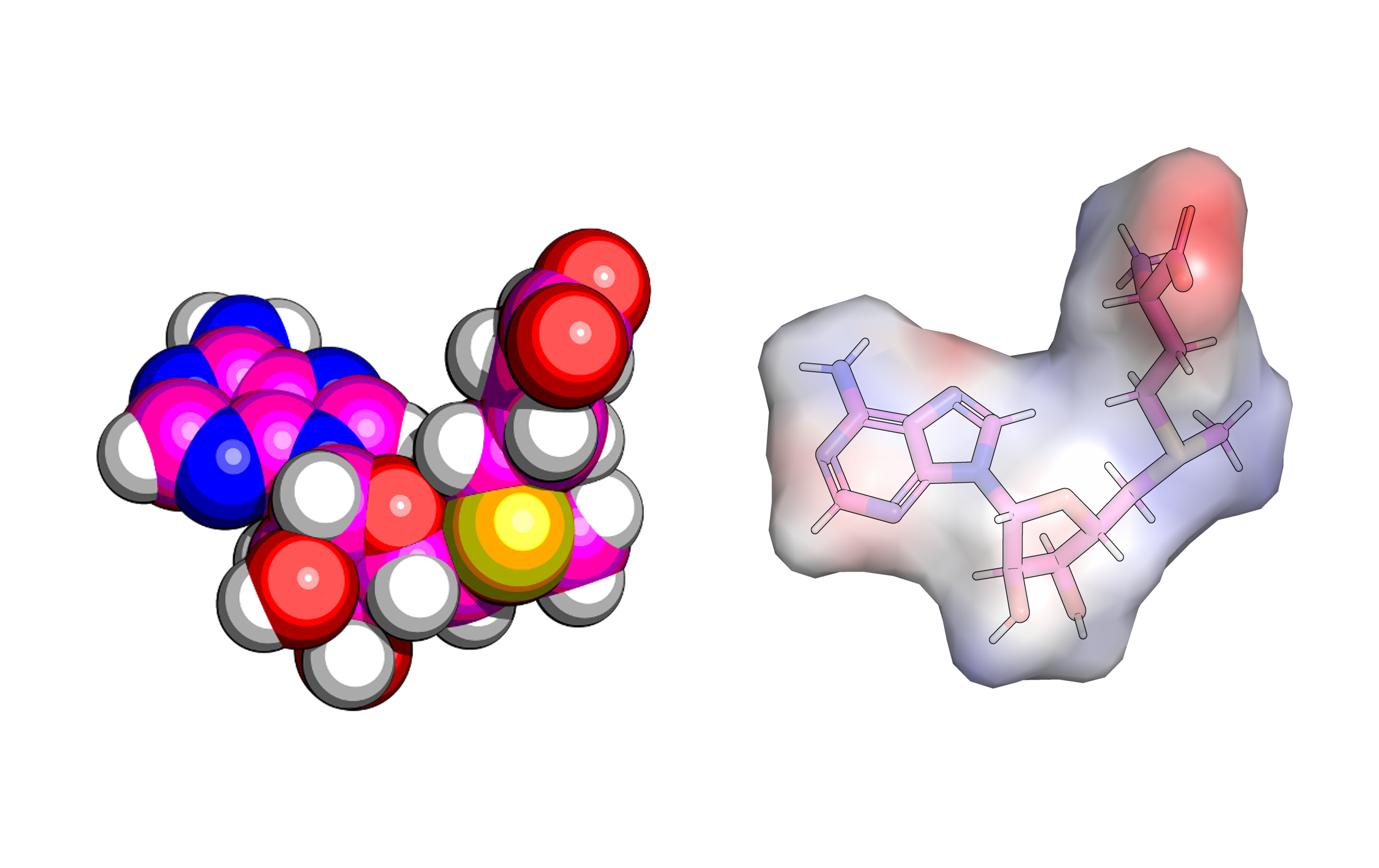
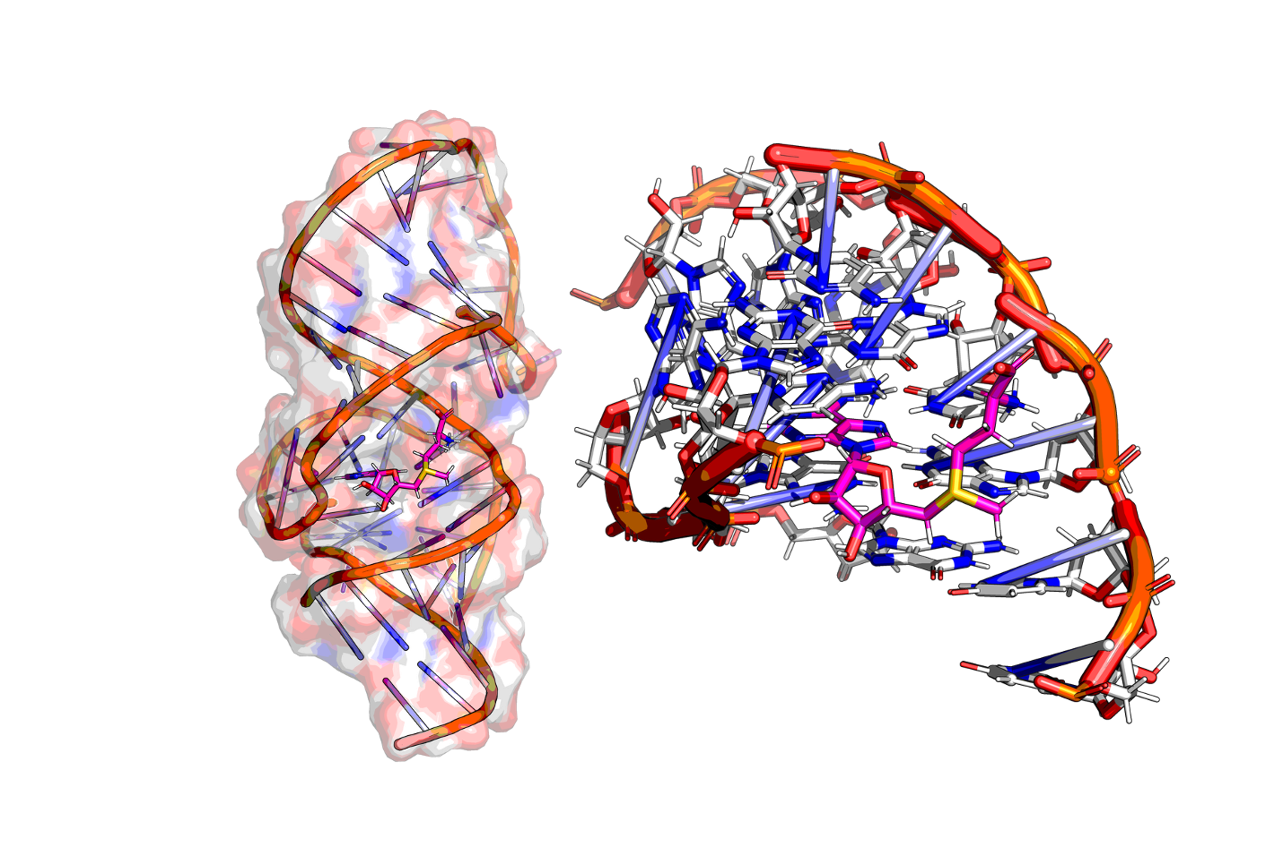
Figure 1. Shape and electrostatic model of S-adenosyl methionine (SAM, top) and SAM-VI-RNA binding model (bottom)
Our methods can account for the unique structural features of RNA, including:
- Conformational flexibility
- Non-canonical base pairing
- Distinct electrostatic environments
Together, these approaches provide detailed structural insight to guide RNA-targeted ligand discovery and optimisation.
Hybrid AI‑Enhanced Virtual Screening
Domainex integrates LBDD and SBDD into a unified, AI‑supported pipeline that:
- Predicts ligand complementarity with high accuracy
- Prioritises selective vs non‑selective binders
- Rapidly ranks our proprietary fragment library for RNA-focused hit discovery
This hybrid approach offers speed, adaptability, and high enrichment, enabling rapid project progression.
Biophysical Assays for Measuring RNA–Small Molecule Binding
Spectral Shift: Medium‑Throughput RNA Binding Analysis
Using labelled RNA, Spectral Shift detects blue/red spectral shifts in the emission spectrum of the fluorophore induced by compound binding.
Strengths of Spectral Shift:
- Medium‑throughput, fast turnaround
- Low RNA consumption
- Highly reproducible, excellent signal‑to‑noise
- Suitable for both hit identification and affinity (KD) determination
- Effective for virtual‑screening follow‑up and fragment screening
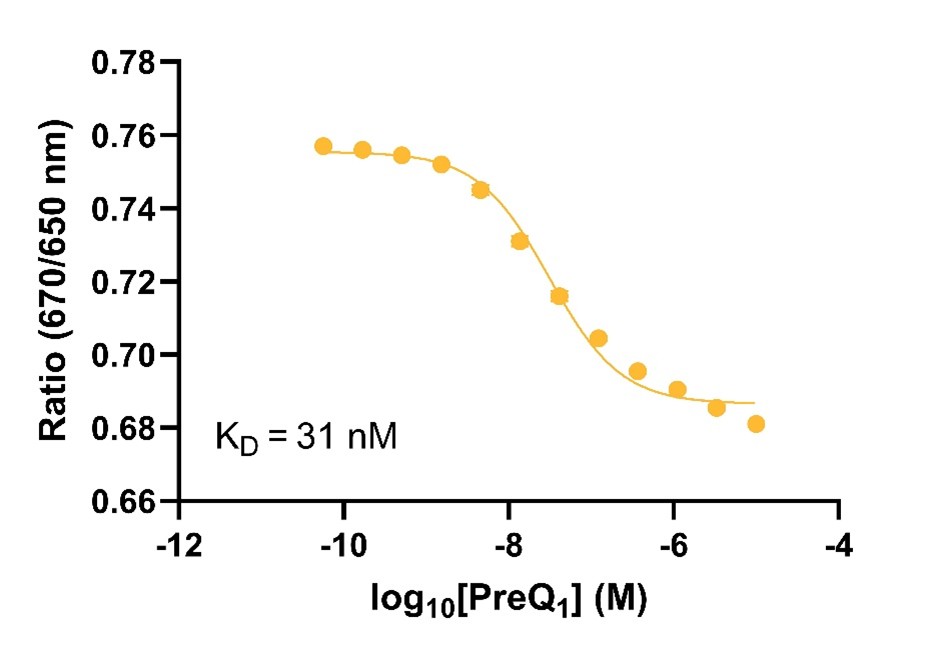
Figure 2: Spectral Shift - Small molecule binding to the Tte preQ1 riboswitch
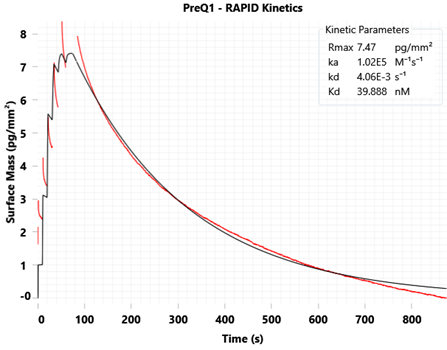
Figure 3: GCI Small molecule binding to the Tte preQ1 riboswitch
Advantages for RNA targets:
- Extremely low RNA consumption
- High sensitivity to small‑molecule binding
- Full kinetic parameter determination (kon, koff)
- Ideal for hit identification, orthogonal validation, and detailed affinity/kinetic analysis
Isothermal Titration Calorimetry (ITC)
Strengths of ITC for RNA targets:
- Complete thermodynamic profiling in a single experiment
- No labelling or immobilization required
- Quantification of folded RNA
- Suitable for the assessment of a wide range of binding affinities and molecules with a wide range of molecular weights
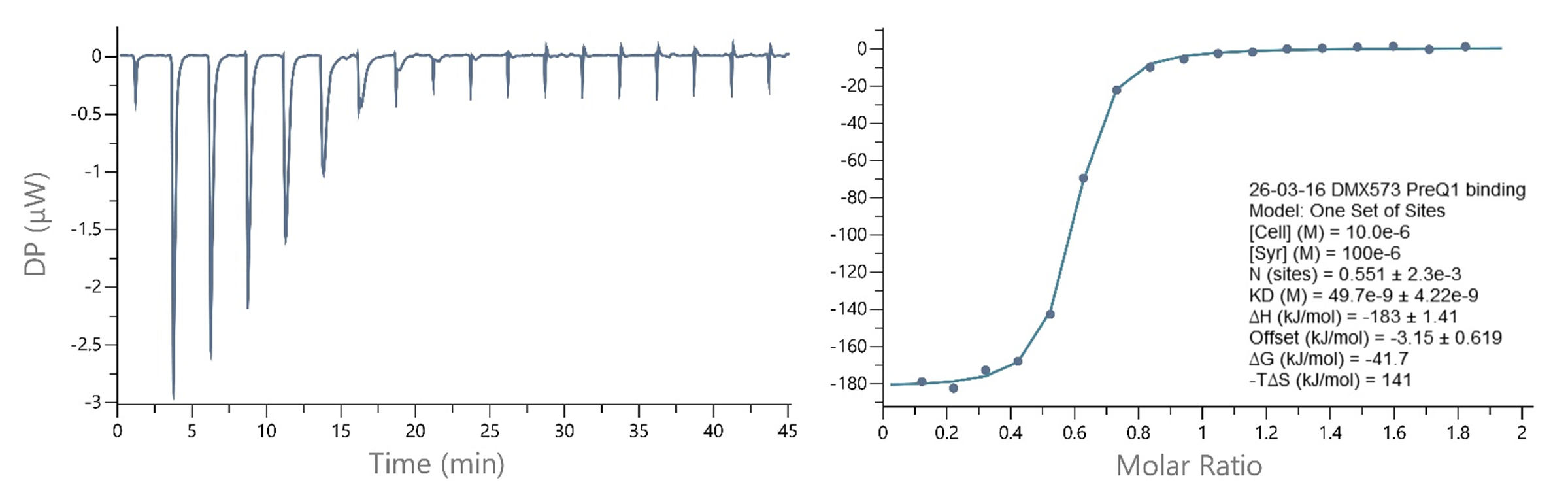
Figure 4: ITC – Small molecule binding to the Tte PreQ1 riboswitch
Post hit generation
Domainex has experience in RNA hit expansion and can apply its direct-to-biology capabilities to significantly shorten the design-make-test cycle and drive RNA drug discovery programs forward at pace.
Frequently Asked Questions
How does Domainex identify RNA binding small molecules?
Domainex offers flexible hit‑finding strategies for RNA targets, using ligand‑based virtual screening, structure‑based modelling, AI‑enhanced hybrid screening, and fragment‑based screening. These approaches can be applied individually or in combination, depending on the target and available data, to efficiently explore RNA‑relevant chemical space and identify druggable RNA pockets. Domainex also maintains a curated library of over 1,000 fragments available for immediate screening as novel starting points.
How do I know which biophysical method is best for my project?
Domainex scientists review your target, modality, and project goals to recommend the most informative and cost‑effective biophysical methods. We regularly design bespoke workflows, selecting the right technique or combination of techniques to answer your specific biological questions.
What biophysical methods does Domainex use to measure RNA–small molecule interactions?
Domainex employs a suite of complementary assays, including:
What is Grating-Coupled Interferometry (GCI)?
GCI is a biosensor-based technique (analogous to SPR) where the target is immobilised onto specialised sensor chips and the passage of analytes over the chip surface are monitored as time-dependent changes in refractive index, which is used to quantify molecular interactions and determine binding kinetics and affinities.
How does GCI differ from SPR?
Where GCI and SPR differ is that SPR detects a localised area of the chip surface whereas GCI samples the entire chip surface. GCI is therefore able to detect more binding events which enhances its sensitivity relative to SPR.
How can RNA be immobilised for GCI/SPR?
Domainex can modify the 5’ or 3’ end with a biotin tag which can be immobilised on streptavidin sensor chips.
What is Spectral Shift?
Spectral Shift is a fluorescence-based biophysical technique that monitors subtle changes in the overall emission profile of the fluorescent tag on your target of interest. Ligand binding typically results in a shift in the emission wavelength through changes in the chemical environment of the fluorophore on the target, either through proximity or conformational changes.
What is Temperature Related Intensity Change (TRIC)?
TRIC is an additional fluorescence-based biophysical technique whereby the fluorescence intensity of a fluorophore changes depending on local temperatures in solution. In addition to changes in fluorescence due to the physical environment, TRIC is strongly influenced by the chemical environment, which can be altered upon the binding of a ligand to the target near the fluorescent label, or by a conformational change of the target as a result of binding to the target elsewhere.
Does Domainex offer MST?
Yes. Domainex offers MST but we typically recommend Spectral Shift or TRIC due to increased throughput and better data quality, especially with Spectral Shift.
How can RNA be labelled for Spectral Shift or MST/TRIC?
The 5’ or 3’ end can be modified with a Cy5 fluorescent tag that can be detected on both NanoTemper instruments.
What is Isothermal Titration Calorimetry (ITC)?
Isothermal Titration Calorimetry (ITC) is a biophysical technique which directly measures the heat released or absorbed during molecular binding events in a label-free environment. It is typically used to study the binding of small molecules to larger macromolecules such as proteins or DNA.
Does Domainex offer high‑throughput or rapid‑turnaround biophysics?
Domainex provides rapid screening for early triage, hit validation, and fragment expansion. For time‑critical projects, we offer accelerated timelines while maintaining data robustness and scientific integrity.
Can Domainex support integrated drug discovery programmes?
The biophysics team at Domainex supports integrated drug discovery programs by working seamlessly with internal medicinal chemistry, structural biology, computational chemistry, and screening groups. This integrated model ensures that biophysical insights directly inform hit identification, hit-to-lead progression, and lead optimisation.
How do I start a project or discuss feasibility?
Contact Domainex through our enquiry form with your target and objectives. Our scientists will arrange a consultation to discuss project scope, feasibility, timelines, and recommended methods, ensuring a smooth and collaborative project launch.
Start your next project with Domainex
Contact one of our experts today